Our growing understanding of extremophiles here on Earth has opened up new possibilities in astrobiology. Scientists are taking another look at resource-poor worlds that appeared like they could never support life. One team of researchers is studying a nutrient-poor region of Mexico to try to understand how organisms thrive in challenging environments.
The researchers worked in a region of Mexico called the Cuatro Ciénegas Basin. About 43 million years ago, the basin was a shallow sea, until it became isolated from the Gulf of Mexico. It's a distinctive region because it's both nutrient-poor and home to aquatic microbes with ancient ancestry.
The lead author of the new study is Jordan Okei from Arizona State University's School of Earth and Space Exploration. The title of the study is " Genomic adaptations in information processing underpin trophic strategy in a whole-ecosystem nutrient enrichment experiment." It's published in the journal eLIFE.
The study focuses on an organism's genome, and fundamental aspects of it like the organism's size, the way that it encodes information, and the density of information. The researchers studied how those characteristics allow an organism to thrive in an extreme environment, like the one at Cuatro Ciénegas Basin. In some ways, the basin is an analog for early Earth, or ancient, wet Mars.
"This area is so poor in nutrients that many of its ecosystems are dominated by microbes and may have similarities to ecosystems from early Earth, as well as to past wetter environments on Mars that may have supported life," said lead author Okie.
There's a cost to everything an organism does, and organisms make many trade-offs as they go about their business. These trade-offs affect the efficiency of an organism's biochemical information processing. An organism that has adapted to and evolved in a nutrient-poor environment may not have "invested" in the capability to use large amounts of resources to replicate itself.
That was the team's hypothesis, and they devised experiments to investigate it.
Associate Professor Christopher Dupont from the J. Craig Venter Institute is a senior author of this study. In a press release, Dupont said "We hypothesized that microorganisms found in oligotrophic (low-nutrient) environments would, out of necessity, rely on low-resource strategies for replication of DNA, transcription of RNA and translation of protein. Conversely, a copiotrophic (high-nutrient) environment favors resource-intensive strategies."
The experiment involved setting up what are called "mesocosms," miniature ecosystems. The organisms were then fed elevated levels of fertilizer containing nitrogen and phosphorous. Those elements spurred increased growth in the microorganisms inside the mesocosms. At the end of the experiment they looked how the community of organisms responded to the increased nutrients, compared to the control groups.
In their study, the authors focused on four traits that govern an organisms ability to process biological information in their cells:
- Multiplicity of genes essential to protein biosynthesis: Copiotrophs, or organisms adapted to nutrient-rich environments, should have higher numbers of genes that contribute to greater growth rates. But there's a trade-off: they're at a disadvantage in nutrient-poor environments, and their higher replication rates could end up reducing their growth efficiency.
- Genome size: An organism with a smaller genome needs fewer resources to replicate, and has a smaller cell size. These organisms can respond more quickly to nutrient-poor conditions after a time of relative nutrient abundance.
- Guanine and cytosine content: Guanine and cytosine are nucleotide bases. Scientists aren't exactly certain why, but organisms with high GC levels in their genome likely do better in resource-rich environments, maybe because GC are more "expensive" to produce. So organisms with lower GC content may do better in resource-poor environments.
- Codon usage bias: Codons are sequences of DNA or RNA nucleotide triplets. Codons specify which amino acid to add next during protein synthesis. Multiple different codons can code an amino acid, but in a nutrient-rich environment, codons that use resources quicker should be biased over their counterparts.
This study is different because it looks at all four of those traits, while previous studies have focused on only one or two of them. This study also looks at how these traits work in a community, whereas previous studies took different approaches. As they say in their paper, "Our study is noteworthy as one of the first whole-ecosystem experiments to involve *experiment-level replicated* metagenomic assessments of community response."
"This study is unique and powerful because it takes ideas from the ecological study of large organisms and applies them to microbial communities in a whole-ecosystem experiment." Senior Author Jim Elser, ASU School of Life Sciences
The experiment lasted 32 days, and took place in Lagunita pond in the Cuatro Ciénegas Basin. During that time the researchers conducted field monitoring, sampling and routine water chemistry.
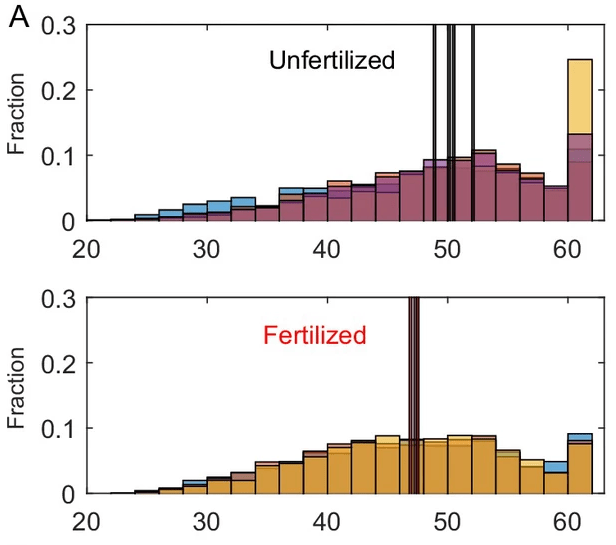
The results were in line with the hypothesis: the mesocosms became dominated by organisms with greater capacity to use the increased nutrients in replication. The control groups were dominated by species that could process biological information at a reduced cost.
"This study is unique and powerful because it takes ideas from the ecological study of large organisms and applies them to microbial communities in a whole-ecosystem experiment," said senior author Jim Elser, from ASU's School of Life Sciences. "By doing so, we were able, perhaps for the first time, to identify and confirm that there are fundamental genomewide traits associated with systematic microbial responses to ecosystem nutrient status, without regard to the species identity of those microbes."
The results of this study tell us something about how life might function in extreme and/or nutrient-poor environments on other worlds. Wherever an organism is, it must have finely-tuned biological information processing capabilities that can take advantage of key resources in their environments. And the environments they find themselves in will determine what those are.
"This is very exciting, as it suggests there are rules of life that should be generally applicable to life on Earth and beyond," said Okie.
 Universe Today
Universe Today